Gene Edit Off Target Analysis Tool - 3 Clicks for a Full Genome Analysis
Built upon BioEngine Technology
Starting at $200.00 per run
The fastest and most thorough Off Target Analysis available, available for wild type CRISPRs. A simple “3 click process”. Enter Reference Sequence, Gene Edit Type and Design and in under one minute receive easy to understand annotated reports with visualizations and all locus information. Primer calculation for off target locations available.
Additional gene edit chemistries / proteins can be analyzed upon request.
Advances in gene editing technology, in particular the CRISPR/Cas9 chemistry, have revolutionized gene editing with promise for advances in basic science, biotechnology, and biomedical research.
With the simplicity of modern gene editing technologies come problematic off target interactions which can lead to failed experiments or cell death.
KBioBox's Gene Edit Off Target Analysis Tool was built to predict potential off targets, allowing scientists to redesign a target, or "know where to look" after a gene edit has been affected. Able to search entire genomes in under a minute, and customizable to any chemistry or enzyme protocol, the Gene Edit Off Target Analysis tool will provide a level of quality assurance previously unavailable for gene edits.
3 Click Submit
Enter cell line, gene edit type, and design, then the Off Target Analysis application built on the BioEngine takes over.
Under a minute
Powered by the BioEngine, searching a whole genome for off-targets takes less than a minute, even for complex chemistries or enzymes
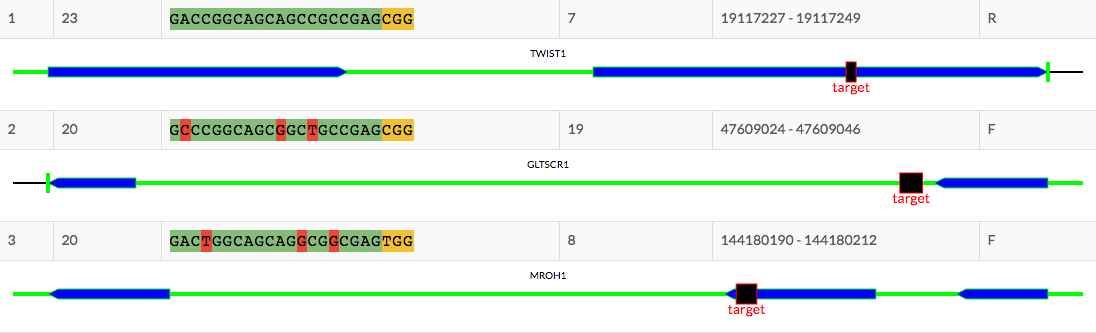
Annotated Reports
Easy to understand reports, with intuitive visualizations and all locus informtaion provided
Learn how to use the online analysis services
Fully Customizable
Unique Enzymes/Chemistries
The Off Target Analysis application can be setup for any enzyme, chemistry, or custom reference.
Custom Reports
In need of additional information, or cross referencing between reports? Reports can be tuned to specific needs
Internal Access
Add the Off Target Analysis application to existing protocols, either through KBioBox's REST based API, or direct integration.
BioEngine Results versus GUIDESeq
Outperforming Open Source Free Tools
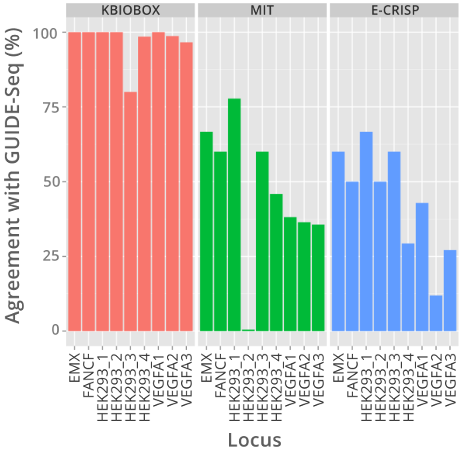
KBioBox's Off Target Analysis application predicted 96.6% of off-targets found by GUIDE-seq, the MIT and E-CRISP tools found less then 50%.
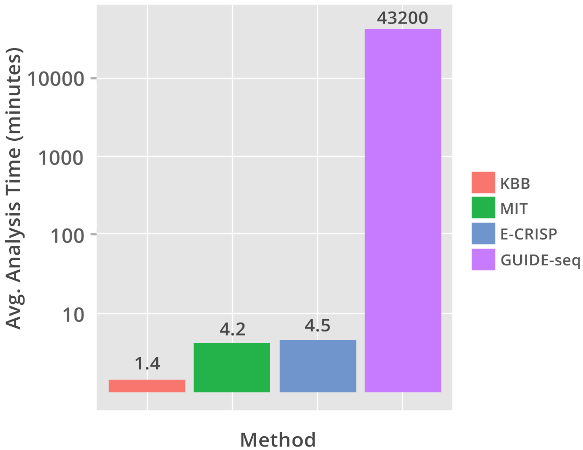
On average KBioBox's Off Target Analysis ran in under two minutes, less than half the time for the MIT and E-CRISP tools.
Built upon KBioBox's BioEngine, the Off-Target Analysis tool predicted 96.6% of off-targets reported by the GUIDE-seq method in under two minutes. By comparison MIT's CRISPR design tool, and the E-CRISP tool both found less than 50% of total off-targets reported by GUIDE-seq.